✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
QuantiTect Whole Transcriptome Kit (100)
Cat. No. / ID: 207045
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Features
- High cDNA yields for unlimited qPCR analysis and archiving
- Equal amplification of all cDNA and all transcript regions
- Fast and easy protocol with optimized reagents
Product Details
The QuantiTect Whole Transcriptome Kit enables the preamplification and reverse transcription of limited amounts of RNA to high yields of cDNA for unlimited gene expression analysis using real-time PCR. An optimized protocol ensures a fast and easy procedure. The kit consists of a complete set of enzymes and buffers for whole transcriptome amplification (WTA). Innovative modification of Phi 29 polymerase technology allows yields of up to 40 μg cDNA from as little as 1 ng RNA. The uniquely high processivity of this polymerase guarantees the generation of cDNA containing uniformly amplified targets to ensure reliable gene expression analysis in real-time PCR.
Performance
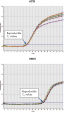
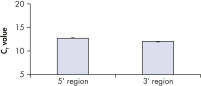
The QuantiTect Whole Transcriptome Kit provides highly uniform amplification of all transcripts, which is essential for reliable gene expression analysis. All mRNA transcripts are amplified with equal representation at both 5' and 3' regions (see figure " Equal amplification of 5' and 3' regions"). Preservation of the transcript profile is demonstrated in the figure " Preservation of transcript profile", where WTA-amplified cDNA prepared using the QuantiTect Whole Transcriptome Kit is compared with nonamplified cDNA prepared using a reverse transcriptase.
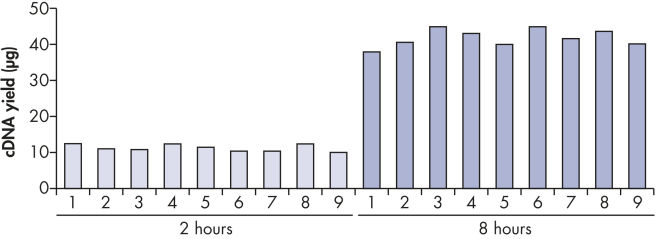
High and reproducible cDNA yields of up to 40 μg can be achieved (see figure " Reproducible cDNA yields"). This allows unlimited real-time PCR analysis with highly reproducible CT values (see table and figure " Reliable real-time PCR analysis"). In addition, since cDNA is more stable than RNA, it can be safely archived for future studies.
| Starting material | Amplification time | Typical yield of amplified product | Number of real-time PCR analyses* |
|---|---|---|---|
| 10 ng RNA | 2 hours | Up to 10 µg cDNA | 1000 |
| 10 ng RNA | 8 hours | Up to 40 µg cDNA | 4000 |
See figures
Principle
Gene expression profiling can be limited by the small amount of biological sample available. By using the QuantiTect Whole Transcriptome Kit, unlimited real-time PCR analyses of small and precious samples are possible. The QuantiTect Whole Transcriptome Kit is optimized for whole transcriptome amplification from as little as 1 ng RNA, or RNA corresponding to about 50 cells. Even lower amounts of RNA can be used, depending on the quality of the RNA and the abundance of the transcript of interest.
The QuantiTect Whole Transcriptome Kit contains reverse transcriptase, DNA polymerase, and optimized buffers and reagents for the amplification of all transcripts within an RNA sample. The kit integrates cDNA synthesis with the proven quality of REPLI-g technology for nonbiased sequence amplification. REPLI-g amplification uses Multiple Displacement Amplification (MDA) technology, which carries out isothermal sequence amplification using a uniquely processive DNA polymerase (see figure " Whole transcriptome amplification"). This technology ensures unbiased amplification of all transcript regions, even of 5' ends, providing unlimited cDNA highly suited for real-time PCR.
See figures
Procedure
See figures
Applications
cDNA prepared using the QuantiTect Whole Transcriptome Kit is intended for use in real-time PCR analysis with QuantiFast or Rotor-Gene Kits and can be archived for future analysis. The cDNA is not suitable for use in microarray analysis. For whole genome amplification from small or precious samples, QIAGEN offers REPLI-g kits and REPLI-g Service. Amplification is highly uniform with minimal sequence bias, providing DNA suitable for applications such as genotyping and Comparative Genome Hybridization (CGH).
Supporting data and figures
Reproducible cDNA yields.
Specifications
| Features | Specifications |
|---|---|
| Amplification | Whole total RNA |
| Starting material | Purified total RNA |
| Reaction time | 5 hours or 11 hours |
| Applications | Real-time PCR, gene expression analysis |
| Minimal pipetting volume needed | 1 µl |
| Samples per run (throughput) | Mid |
| Technology | Reverse transcription followed by ligation and multiple displacement amplification (MDA) |
| Yield | 10 µg or 40 µg cDNA |
| Maximum input volume | 5 µl total RNA |
| Starting amount of total RNA | >10 ng purified total RNA |
| Reaction volume | 50 µl |