Real-time PCR을 이용하여 Hepatitis B virus DNA를 정량 검사할 수 있습니다
Products
Features
- CE-marked — full compliance with European IVD Directive 98/79/EC
- 신뢰할 수 있는 결과 — DNA 분리 과정과 PCR 과정이 잘 수행되었는지 증명하기 위한 internal control 이용
- 고감도 검출 — 0.01 IU/µl (artus HBV TM PCR Kit와 ABI 7700 SDS 장비를 이용하여 검증)
- Broad linear range — from 0.02 IU/µl to 1 x 108 IU/µl (valid for the artus HBV RG PCR Kit on the Rotor-Gene 3000)
- 제공되는 5개의 standard를 이용하여 HBV DNA를 정확하게 정량할 수 있음
Product Details
Performance
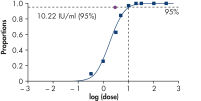
To ensure highest sensitivity, artus HBV Kits have been optimized to detect low numbers of HBV DNA. The analytical sensitivity of the artus HBV QS-RGQ Kit is 10.2 IU/ml in consideration of the purification and assay setup using the QIAsymphony RGQ system (see figure " Highly sensitive detection of HBV DNA"). (1 IU/ml corresponds to 8.21 copies/ml for detection of HBV DNA on the Rotor-Gene Q. The conversion factor is an approximation based on an average factor across the assay's dynamic range.)
For highest specificity, validation of the artus HBV Kits was carried out using various HBV isolates, including all genotypes A–H and related pathogens.
| Kit | artus HBV RG PCR Kit | artus HBV QS-RGQ Kit |
|---|---|---|
| Validated sample type | EDTA plasma | EDTA plasma |
| Analytical sensitivity | 3.8 IU/ml | 10.2 IU/ml |
| Linear range | 1.1 to >4 x 109 IU/ml | 31.6 to >2 x 107 IU/ml |
| Specificity | HBV genomes A–H | HBV genomes A–H |
See figures
Principle
artus HBV Kits are based on the amplification and simultaneous detection of a specific region of the HBV genome using real-time PCR. The kits provide high levels of specificity, sensitivity (see figure " Highly sensitive detection of HBV DNA"), and reproducibility over a broad linear range.
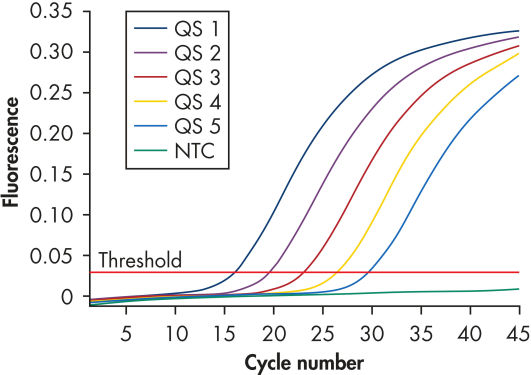
Each artus HBV Kit provides 5 HBV quantitation standards (see figure " Reliable quantitation of HBV load"). Use of the standards enables accurate quantitation of viral load. In addition, the kits contains a second heterologous amplification system to identify possible PCR inhibition. This is detected as an internal control (IC) in a different fluorescence channel from the analytical PCR. The detection limit of the analytical HBV PCR is not reduced.
| Kit | artus HBV RG PCR Kit and artus HBV QS-RGQ Kit |
|---|---|
| Validated sample type | EDTA plasma |
| Amplicon | 134 bp core region of the HBV genome |
See figures
Procedure
artus HBV PCR Kits provide all necessary reagents optimized for reliable HBV DNA detection and quantitation. Simply add template DNA to the ready-to-use PCR master mix (and Mg solution, LC kit only), and start the reaction on the appropriate real-time cycler using the optimized cycling program described in the kit handbook.
Complete automated system from sample to HBV detection
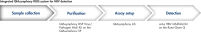
The QIAsymphony RGQ workflow solution for HBV detection comprises the QIAsymphony SP for sample preparation, the QIAsymphony AS for assay setup, and the artus HBV QS-RGQ Kit on the Rotor-Gene Q. The system enables reliable pathogen detection with a complete CE-IVD-compliant workflow (see figure "Integrated QIAsymphony RGQ system for HBV detection").
Recommendations for manual viral RNA purification
artus HBV LC, RG, and TM PCR Kits are validated for use with viral RNA purified from EDTA plasma using the CE-marked QIAamp DSP Virus Kit.
Applications
artus HBV LC, RG, and TM PCR Kits enable rapid and sensitive detection and quantitation of HBV DNA purified from human plasma using the QIAamp DSP Virus Kit. Kits are available for use on the ABI PRISM 7000, 7700, and 7900HT SDS, on the LightCycler 1.1/1.2/1.5/2.0 Instruments, or on Rotor-Gene Q instruments.
The artus HBV QS-RGQ Kit is designed to be used with the QIAsymphony RGQ system, providing a complete CE-IVD-compliant workflow from sample to HBV DNA detection and quantitation.
Supporting data and figures
Reliable quantitation of HBV load.