✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
QIAseq FastSelect –rRNA Fish Kit (8)
Cat. No. / ID: 333251
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Features
- Compatible with QIAGEN, Illumina, NEB, KAPA and other RNA-seq library kits
- High-performance cytoplasmic and mitochondrial rRNA removal in just 14 minutes
- Only one pipetting step – combine QIAseq FastSelect reagent with RNA and incubate
- No extra cleanup steps or NGS library protocol changes
- Designed to cover Danio rerio (zebrafish) and Salmo salar (salmon) and related species
Product Details
QIAseq FastSelect –rRNA Fish Kits use a novel method to remove highly abundant RNA that is of low scientific value from your RNA-seq libraries. Researchers using RNA-seq for whole transcriptome analysis can use QIAseq FastSelect –rRNA Fish Kits to remove all annotated rRNAs in RefSeq and Ensembl for Danio rerio (zebrafish) and Salmo salar (atlantic salmon). Depending on rRNA sequence homology, QIAseq FastSelect −rRNA Fish will work on additional fish species (dependent upon target rRNA sequence homology with D. rerio or S. salar).
Analyze RNA-seq data with ease using the GeneGlobe-integrated RNA-seq Analysis Portal – an intuitive, web-based data analysis solution created for biologists and included with this QIAseq Kit.
We recommend using QIAseq Stranded RNA Library Kits for robust strand-specific RNA-seq library preparation for high-quality and highly fragmented (FFPE) RNA. QIAseq Stranded RNA Library Kits utilize unique dual indexing (UDIs) which enables multiplexing of up to 384 samples per flow cell lane on Illumina NGS instruments and mitigates index hopping on Illumina NovaSeq and Illumina patterned flow cells.
Design your own custom QIAseq FastSelect pools to remove any RNAs you wish from your RNA-seq library – take a look at our QIAseq FastSelect Custom RNA Removal Kits.
Want to try the QIAseq FastSelect –rRNA Fish Kit for the first time? Request a trial kit to evaluate.
Principle
Removing highly expressed, but biologically unimportant RNA transcripts makes NGS more efficient and enables higher sample throughput with higher sensitivity. Furthermore, removal of unwanted RNA species from full length and fragmented RNA samples can be particularly challenging and can result in suboptimal performance.
QIAseq FastSelect –rRNA Fish Kits are designed for quick, efficient removal of cytoplasmic and mitochondrial fish ribosomal RNA from total RNA during NGS RNA library preparation. QIAseq FastSelect seamlessly integrates with your existing RNA stranded library preparation workflow for RNA removal in a single, 14-minute inline step. Prior to RNA heat fragmentation (which is optional and dependent upon the library preparation kit and sample type), QIAseq FastSelect removal reagent is directly combined with total RNA and the library preparation-specific buffers. After fragmentation, the reaction temperature is stepwise cooled to room temperature and the remaining library preparation steps are completed. There is no need to perform any type of enrichment on the total RNA samples. QIAseq FastSelect –rRNA Fish Kits ensure consistently high performance with RNA amounts ranging from as little as 1 ng up to 1 μg. QIAseq FastSelect can be used with RNA from fresh samples from many species of fish (dependent upon target rRNA sequence homology with D. rerio or S. salar), and delivers reliable rRNA removal and high reproducibility in downstream applications.
Procedure
Most RNA removal or depletion strategies associated with RNA-seq library construction are sample pre-treatment strategies involving hybrid-capture or enzymatic removal of unwanted RNA. Our unique QIAseq FastSelect procedure is compatible with QIAGEN, Illumina, KAPA and NEB and other RNA library kits and provides complete rRNA removal in a single, 14-minute inline step (see figure QIAseq FastSelect Kit workflow ). This is dramatically faster than alternative RNA depletion kits, which require pre-treatment protocols involving more than 25 steps and 2 hours to complete.
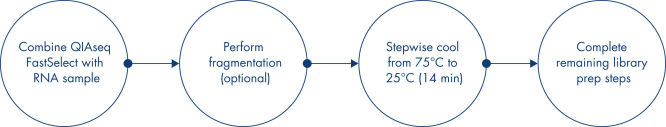
Simply add QIAseq FastSelect reagent to the RNA sample, perform fragmentation (if required), stepwise cool the reaction from 75°C to 25°C for 14 minutes and then complete the remaining library preparation steps. QIAseq FastSelect works with or without RNA fragmentation, providing the flexibility to use fragmented RNA from fish samples or high-quality RNA as part of a standard RNA-seq library construction workflow.
See figures
Applications
QIAseq FastSelect delivers rapid, reliable RNA removal from full length or fragmented RNA samples from many species of fish (dependent upon target rRNA sequence homology with D. rerio or S. salar). QIAseq FastSelect –rRNA Fish Kits are available in a variety of different formats and sizes to suit your specific applications.
Supporting data and figures
QIAseq FastSelect Kit workflow
Simply add QIAseq FastSelect reagent (rRNA Removal and/or Globin Removal) to your RNA sample, perform fragmentation (if required), then initiate a stepwise cool-down from 75°C to 25°C over 14 minutes. Researchers can then complete the remaining library preparation steps without any additional changes to their workflow. QIAseq FastSelect works with or without RNA fragmentation, providing the flexibility to use FFPE or degraded RNA samples, or high-quality RNA as part of a standard RNA-seq library construction workflow.
QIAseq FastSelect –rRNA HMR Kits, QIAseq FastSelect –Globin Kits, QIAseq FastSelect –rRNA/Globin Kit, QIAseq FastSelect –rRNA Plant Kits, QIAseq FastSelect –rRNA Yeast Kits, QIAseq FastSelect –rRNA Worm Kits, QIAseq FastSelect –rRNA Fly Kits, QIAseq FastSelect –rRNA Fish Kits and QIAseq FastSelect Epidemiology Kits have been tested with the QIAseq Stranded Total RNA Lib Kit (QIAGEN), TruSeq Stranded (Illumina), NEBNext Ultra II Directional (New England Biolabs) and KAPA RNA HyperPrep (KAPA Biosystems) library preparation kits.