REPLI-g FFPE Kit
直接对福尔马林固定、石蜡包埋组织(FFPE)中的DNA进行全基因组扩增
直接对福尔马林固定、石蜡包埋组织(FFPE)中的DNA进行全基因组扩增
✓ 全天候自动处理在线订单
✓ 博学专业的产品和技术支持
✓ 快速可靠的(再)订购
Cat. No. / ID: 150243
✓ 全天候自动处理在线订单
✓ 博学专业的产品和技术支持
✓ 快速可靠的(再)订购
如果基因组DNA量不足以进行基因组分析,这一问题可通过对样本内全基因组DNA扩增解决。REPLI-g FFPE Kit包含DNA聚合酶、新型缓冲液和试剂,确保对福尔马林固定、石蜡包埋(FFPE)组织进行高效的全基因组扩增,无需预先纯化DNA。组织切片裂解后,对DNA进行处理后进行DNA片段连接,然后使用REPLI-g技术扩增长链DNA。
提供两种实验方案,扩增时间不同。标准2小时扩增反应方案,每50 µl反应体系的常规产量为10 µg,高产量反应(8小时扩增)可得到40 µg产物。
用于基因分型研究的基因组DNA量通常不足。全基因组扩增(WGA)通过在样本内扩增全基因组,克服了这个难题,可扩增获得足量的来源于同一样本的DNA用于后续分析。
对福尔马林固定、石蜡包埋的组织样本(FFPE)进行基因分型,可将组织的形态变化直接与特定基因组变异联系起来。但是,福尔马林固定会导致DNA片段化,并与样本内的其他生物分子发生交联。此外,从FFPE样本中纯化得到的DNA量较少,只能用于少数几个分析。
REPLI-g FFPE Kit利用新型的DNA处理技术,将DNA片段连接起来(参见" REPLI-g FFPE principle")。然后通过经验证的REPLI-g技术对随机连接起来的DNA进行全基因组扩增。REPLI-g产品应与MDA以及独特的DNA聚合酶结合使用。REPLI-g DNA Polymerase凭借其活跃的进行性酶反应和单链置换反应,可很大程度上减少序列偏差和不均一位点的出现。该方法得到的结果比基于PCR的WGA方法更加可靠。新型的DNA处理技术可将MDA技术充分利用在FFPE组织样本中高度降解的DNA上。REPLI-g FFPE Kit在位点不变化的同时,实现对珍贵样本材料中DNA的扩增,产物可用于多种下游应用。
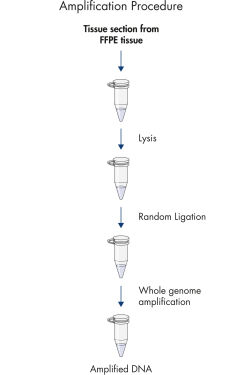
REPLI-g FFPE可使用FFPE组织样本直接进行DNA扩增,无需预先纯化DNA。裂解组织切片后,使用新型缓冲液和酶处理DNA,将片段化的DNA连接起来(参见" REPLI-g FFPE procedure"和" High-molecular-weight ligated DNA")。反应后生成DNA长链,长链经REPLI-g技术扩增。扩增得到的DNA可即用于多数基因分型下游应用,无需进行纯化。
使用REPLI-g FFPE Kit扩增得到的DNA适用于real-time PCR(如使用QuantiFast Kits)和终点式PCR(如使用QIAGEN Fast Cycling PCR Kit)。PCR扩增子应比起始模板的平均片段长度短。还适用于微卫星分析(参见" Reliable microsatellite analysis")和SNP基因分型(REPLI-g FFPE扩增得到的DNA可能不适用于需要限制性内切进行DNA标记的基因分型应用)。
REPLI-g FFPE扩增的DNA适合以下应用:
| Features | Specifications |
|---|---|
| Yield | 10 µg (2h reaction)-40 µg (8h reaction) |
| Quality assessment | No |
| Starting material | FFPE tissue sections or purified genomic DNA |
| Technology | MDA |
| Maximum input volume | 1 tissue section (10-40 µm thickness) |
| Reaction time | 2-8h (2h: standard; 8h: high-yield reaction) |
| Reaction volume | 50 µl |
| Starting amount of DNA | 100-300 ng* |
| ApplicationsZH | Genotyping, PCR, Real-time PCR |
| Samples per run (throughput) | medium |
| Amplification | Whole genomic DNA |